Research study tests Environmental DNA (eDNA) approaches to detecting arboreal mammals in the EcoPreserve
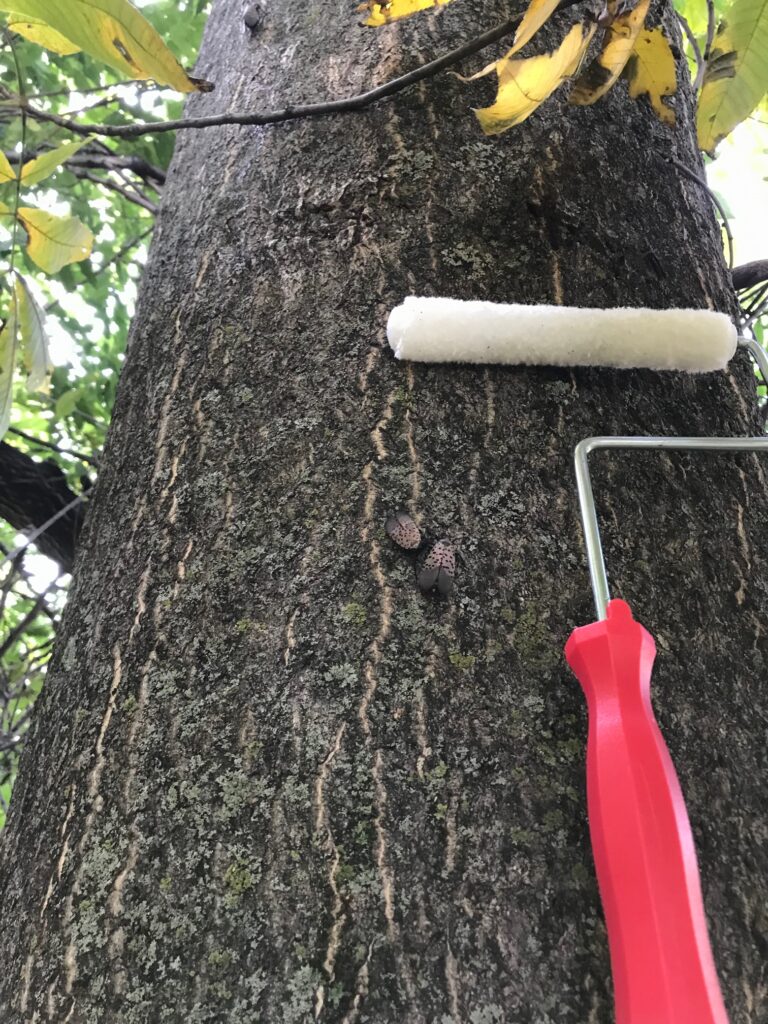

Researchers swab the trunk of an ailanthus tree to then bring back to the lab to analyze for traces of animal DNA
A team from Dr. Julie Lockwood’s research lab used the EcoPreserve and Jockey Hollow National Historical Park to test the efficacy of sampling environmental DNA from trees and soil to detect cryptic arboreal mammals. The eDNA results form the EcoPreserve compared favorably with the Snapshot USA wildlife cam monitoring conducted by Dr. Rick Lathrop’s and bat surveys conducted by Dr. Brooke Maslos’s research teams.
The article with Dr. Mike Allen as first author was published in the January 2023 issue of Nature Scientific reports https://www.nature.com/articles/s41598-023-27512-8
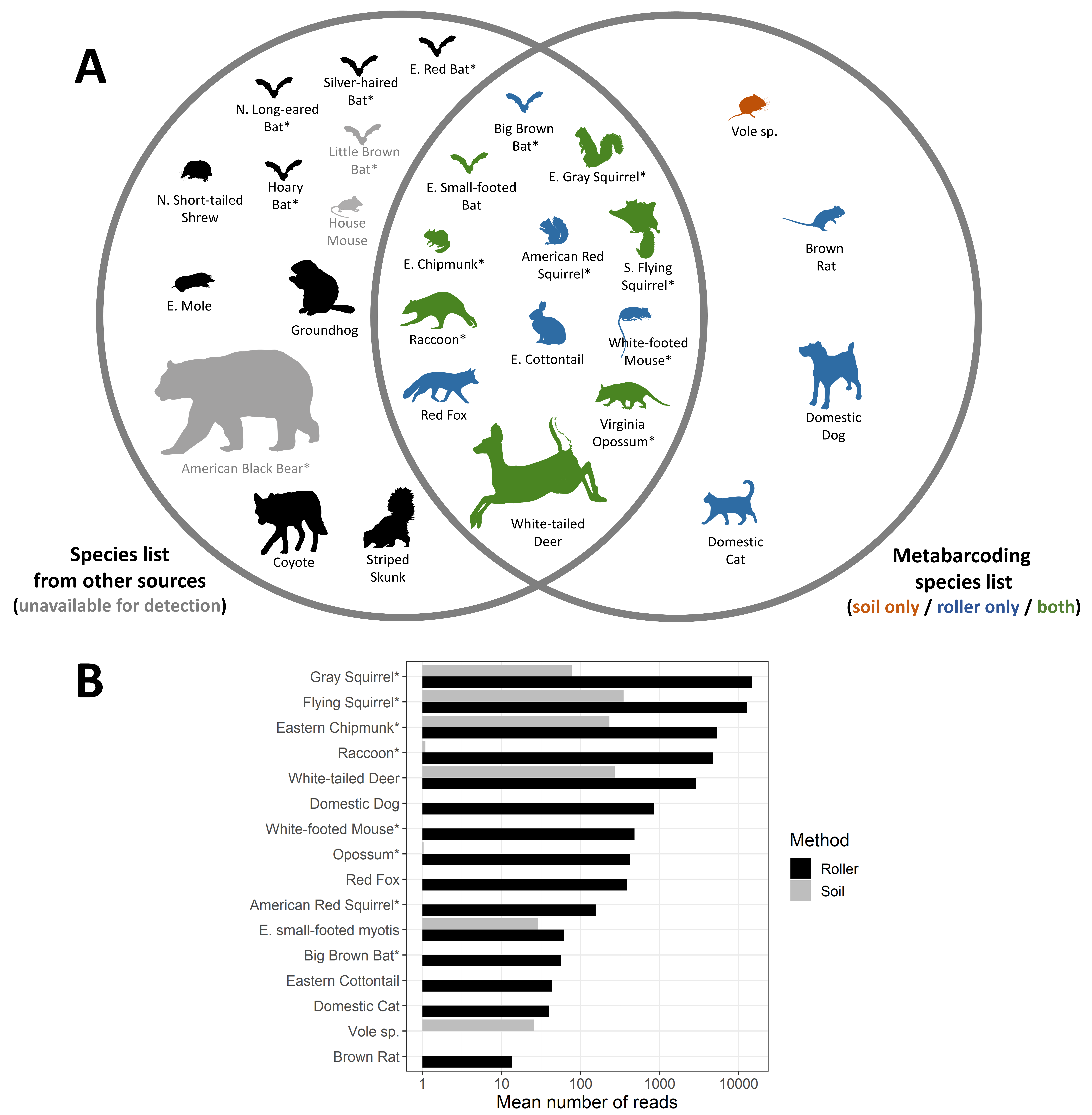
The graphic below (from the paper cited above) depicts the mammal species detected with eDNA metabarcoding using soil and tree bark surface (‘roller’) sampling in two woodlands in central New Jersey, USA. Venn diagram (A) shows which species were detected with metabarcoding (right circle) and which were previously reported as present in the parks. Color coding: blue—detected with metabarcoding of roller samples only; orange—detected with metabarcoding of soil samples only; green—detected with metabarcoding of both roller and soil samples; black—not detected using metabarcoding; gray—species deemed unavailable for metabarcoding detection due to contamination (mouse and bat) or poor performance of the primer set. B) shows the mean number of metabarcoding reads per sample for each species detected. Arboreal species are identified with an asterisk (*).